Inorganic Pyrophosphatase
(All numbering and residues are taken from first PDB file)
![]()
![]()
Bending Residue Dihedral Analysis
Residue
iResidue
i+1Distance of hinge axis to residue i in
(A) Distance of hinge axis to residue i in
(A) Change in
(deg) Change in
(deg) Angle of psi(i) axis to hinge axis
(deg) Angle of psi(i) axis to hinge axis
(deg) Percentage Progress
ASN-18
SER-19
4.8
4.5
7.1
-9.5
101.9
104.1
23.0
SER-19
MET-20
1.5
1.4
4.1
0.5
47.1
47.1
15.7
MET-20
PRO-21
2.8
2.9
-6.0
14.3
83.0
82.9
-22.9
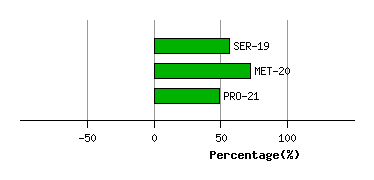
Graph shows rotational transition at bending residues and can be used
to identify hinge bending residues.
Probably only informative for interdomain rotations greater than 20 degrees
Residue
iResidue
i+1Distance of hinge axis to residue i in
(A) Distance of hinge axis to residue i in
(A) Change in
(deg) Change in
(deg) Angle of psi(i) axis to hinge axis
(deg) Angle of psi(i) axis to hinge axis
(deg) Percentage Progress
ASN-84
SER-85
7.5
7.2
12.2
-4.0
50.8
46.1
80.0
SER-85
TYR-86
9.3
9.0
8.4
-12.3
98.2
93.4
1.3
TYR-86
PRO-87
7.8
7.2
-18.5
3.6
121.3
119.7
-80.4
PRO-87
GLU-88
5.5
5.2
-25.4
10.2
64.3
54.4
188.6
GLU-88
ALA-89
1.8
1.6
4.6
1.8
133.9
130.6
-30.0
ALA-89
GLU-90
2.4
2.2
6.5
-15.6
72.4
73.3
-31.8
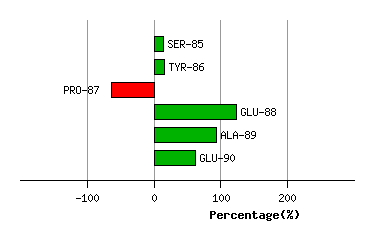
Graph shows rotational transition at bending residues and can be used
to identify hinge bending residues.
Probably only informative for interdomain rotations greater than 20 degrees