Phosphoglycerate Kinase
(All numbering and residues are taken from first PDB file)
![]()
![]()
Bending Residue Dihedral Analysis
Residue
iResidue
i+1Distance of hinge axis to residue i in
(A) Distance of hinge axis to residue i in
(A) Change in
(deg) Change in
(deg) Angle of psi(i) axis to hinge axis
(deg) Angle of psi(i) axis to hinge axis
(deg) Percentage Progress
LEU-178
SER-179
2.1
2.8
-2.9
-4.4
34.4
25.9
39.2
SER-179
LYS-180
1.6
2.3
-5.7
-0.6
86.1
90.1
14.4
LYS-180
VAL-181
2.3
1.5
3.4
3.8
63.6
63.5
42.7
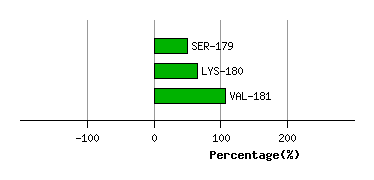
Graph shows rotational transition at bending residues and can be used
to identify hinge bending residues.
Probably only informative for interdomain rotations greater than 20 degrees
Residue
iResidue
i+1Distance of hinge axis to residue i in
(A) Distance of hinge axis to residue i in
(A) Change in
(deg) Change in
(deg) Angle of psi(i) axis to hinge axis
(deg) Angle of psi(i) axis to hinge axis
(deg) Percentage Progress
ALA-196
LYS-197
14.0
14.8
-0.4
-66.5
134.7
110.1
-152.9
LYS-197
VAL-198
15.6
17.4
71.5
-5.6
60.5
34.1
209.7
VAL-198
SER-199
17.2
18.2
30.6
-12.3
36.3
41.5
60.0
SER-199
ASP-200
20.2
20.3
-8.8
17.0
151.2
120.5
37.2
ASP-200
LYS-201
17.9
17.5
19.7
-23.0
64.4
52.2
-0.3
LYS-201
ILE-202
15.2
15.5
38.4
-10.0
76.6
46.8
66.7
ILE-202
GLY-203
17.8
18.3
-11.8
3.3
161.6
161.5
-50.4
GLY-203
VAL-204
18.8
17.1
-6.8
12.4
129.0
98.4
31.0
VAL-204
ILE-205
15.5
13.6
-31.8
-2.0
87.8
100.5
11.7
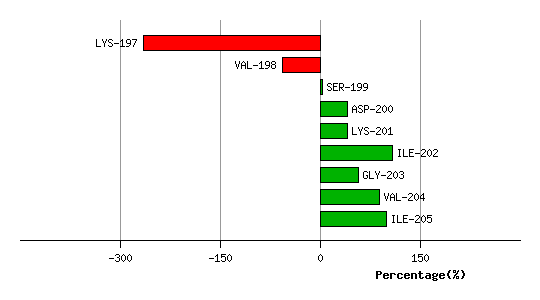
Graph shows rotational transition at bending residues and can be used
to identify hinge bending residues.
Probably only informative for interdomain rotations greater than 20 degrees
Residue
iResidue
i+1Distance of hinge axis to residue i in
(A) Distance of hinge axis to residue i in
(A) Change in
(deg) Change in
(deg) Angle of psi(i) axis to hinge axis
(deg) Angle of psi(i) axis to hinge axis
(deg) Percentage Progress
LEU-208
MET-209
13.7
12.7
8.9
7.2
92.6
84.0
60.1
MET-209
GLU-210
16.8
15.9
-20.2
-3.8
27.6
43.1
70.4
GLU-210
LYS-211
16.6
16.2
28.9
5.5
100.3
102.2
-15.4
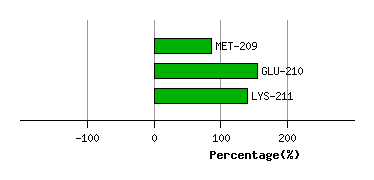
Graph shows rotational transition at bending residues and can be used
to identify hinge bending residues.
Probably only informative for interdomain rotations greater than 20 degrees
Residue
iResidue
i+1Distance of hinge axis to residue i in
(A) Distance of hinge axis to residue i in
(A) Change in
(deg) Change in
(deg) Angle of psi(i) axis to hinge axis
(deg) Angle of psi(i) axis to hinge axis
(deg) Percentage Progress
VAL-350
VAL-351
1.0
1.4
-4.0
-1.5
35.5
41.7
5.1
VAL-351
GLY-352
1.4
1.1
-0.3
47.7
109.1
105.4
56.3
GLY-352
GLY-353
5.0
4.7
-78.8
-65.8
42.9
50.6
420.3
GLY-353
GLY-354
7.9
6.8
55.6
117.4
127.4
150.0
-788.2
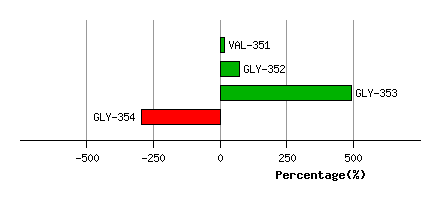
Graph shows rotational transition at bending residues and can be used
to identify hinge bending residues.
Probably only informative for interdomain rotations greater than 20 degrees
Residue
iResidue
i+1Distance of hinge axis to residue i in
(A) Distance of hinge axis to residue i in
(A) Change in
(deg) Change in
(deg) Angle of psi(i) axis to hinge axis
(deg) Angle of psi(i) axis to hinge axis
(deg) Percentage Progress
SER-370
HIS-371
6.5
7.1
-2.7
-2.5
100.5
95.6
-10.1
HIS-371
VAL-372
4.8
5.2
-7.3
0.3
40.3
39.6
22.5
VAL-372
SER-373
5.1
5.1
-20.9
69.2
110.4
108.7
72.1
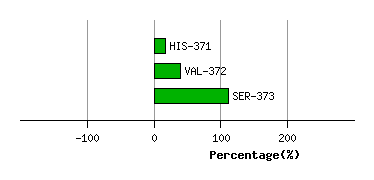
Graph shows rotational transition at bending residues and can be used
to identify hinge bending residues.
Probably only informative for interdomain rotations greater than 20 degrees