Nadp Isocitrate Dehydrogenase
(All numbering and residues are taken from first PDB file)
![]()
![]()
Bending Residue Dihedral Analysis
Residue
iResidue
i+1Distance of hinge axis to residue i in
(A) Distance of hinge axis to residue i in
(A) Change in
(deg) Change in
(deg) Angle of psi(i) axis to hinge axis
(deg) Angle of psi(i) axis to hinge axis
(deg) Percentage Progress
ARG-137
GLU-138
3.9
3.9
0.9
1.2
36.4
37.2
19.4
GLU-138
GLY-139
1.8
1.9
-2.2
3.0
75.5
76.6
4.5
GLY-139
ASN-140
3.1
3.0
-7.9
-6.9
45.7
48.3
124.3
ASN-140
SER-141
5.9
5.7
10.2
-5.1
156.6
156.0
-120.3
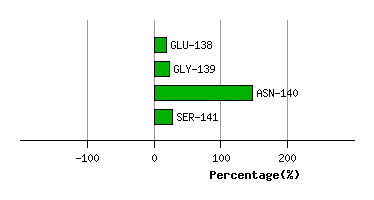
Graph shows rotational transition at bending residues and can be used
to identify hinge bending residues.
Probably only informative for interdomain rotations greater than 20 degrees
Residue
iResidue
i+1Distance of hinge axis to residue i in
(A) Distance of hinge axis to residue i in
(A) Change in
(deg) Change in
(deg) Angle of psi(i) axis to hinge axis
(deg) Angle of psi(i) axis to hinge axis
(deg) Percentage Progress
MET-561
LEU-562
1.6
1.4
-5.8
7.0
49.8
48.2
32.8
LEU-562
SER-563
2.6
2.4
7.0
-8.1
144.9
141.7
18.6
SER-563
VAL-564
1.0
1.0
0.7
-2.9
146.2
143.6
30.2
VAL-564
VAL-565
2.6
2.6
-8.1
10.0
34.3
35.9
23.1
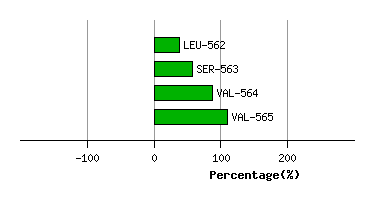
Graph shows rotational transition at bending residues and can be used
to identify hinge bending residues.
Probably only informative for interdomain rotations greater than 20 degrees